Watch the World’s Smallest Data Recorder
S. Himmelstein | November 26, 2017An ordinary laboratory strain of a microbe commonly found in the human gastrointestinal tract — Escherichia coli — has been modified to function as a microscopic data recorder. The bacterial system could be used for environmental monitoring and disease diagnosis, according to Columbia University Medical Center researchers.
CRISPR-Cas, an immune system in many species of bacteria, is tapped to copy snippets of DNA from invading 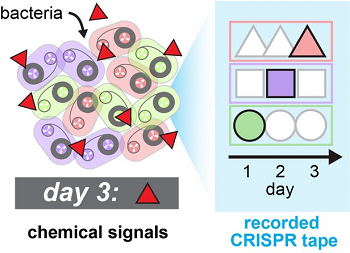 This is the first time CRISPR has been used to record cellular activity and the timing of those events. Source: Wang Lab/Columbia University Medical Centerviruses so that subsequent generations of bacteria can repel these pathogens more effectively. The CRISPR locus of the bacterial genome accumulates a chronological record of the bacterial viruses that it and its ancestors have survived. When those same infectious viruses are encountered, the CRISPR-Cas system can recognize and eliminate them.
This is the first time CRISPR has been used to record cellular activity and the timing of those events. Source: Wang Lab/Columbia University Medical Centerviruses so that subsequent generations of bacteria can repel these pathogens more effectively. The CRISPR locus of the bacterial genome accumulates a chronological record of the bacterial viruses that it and its ancestors have survived. When those same infectious viruses are encountered, the CRISPR-Cas system can recognize and eliminate them.
“When you think about recording temporally changing signals with electronics, or an audio recording…that’s a very powerful technology, but we were thinking how can you scale this to living cells themselves?” said researcher Ravi Sheth.
A piece of DNA called a plasmid was modified to provide it the ability to create more copies of itself in the bacterial cell in response to an external signal. A separate recording plasmid, which drives the recorder and marks time, expresses components of the CRISPR-Cas system. In the absence of an external signal, only the recording plasmid is active, and the cell adds copies of a spacer sequence to the CRISPR locus in its genome. When an external signal is detected by the cell, the other plasmid is also activated, leading to insertion of its sequences instead. The result is a mixture of background sequences that record time and signal sequences that change depending on the cell’s environment.
The researchers examine the bacterial CRISPR locus and use computational tools to read the recording and its timing. The system was demonstrated to handle at least three simultaneous signals and records for days.
This is the first time CRISPR has been used to record cellular activity and the timing of those events. The technology was previously used by synthetic biologists to store poems, books and images in DNA.




